ENCGM023HNS
Summary
- Status
- released
- Description
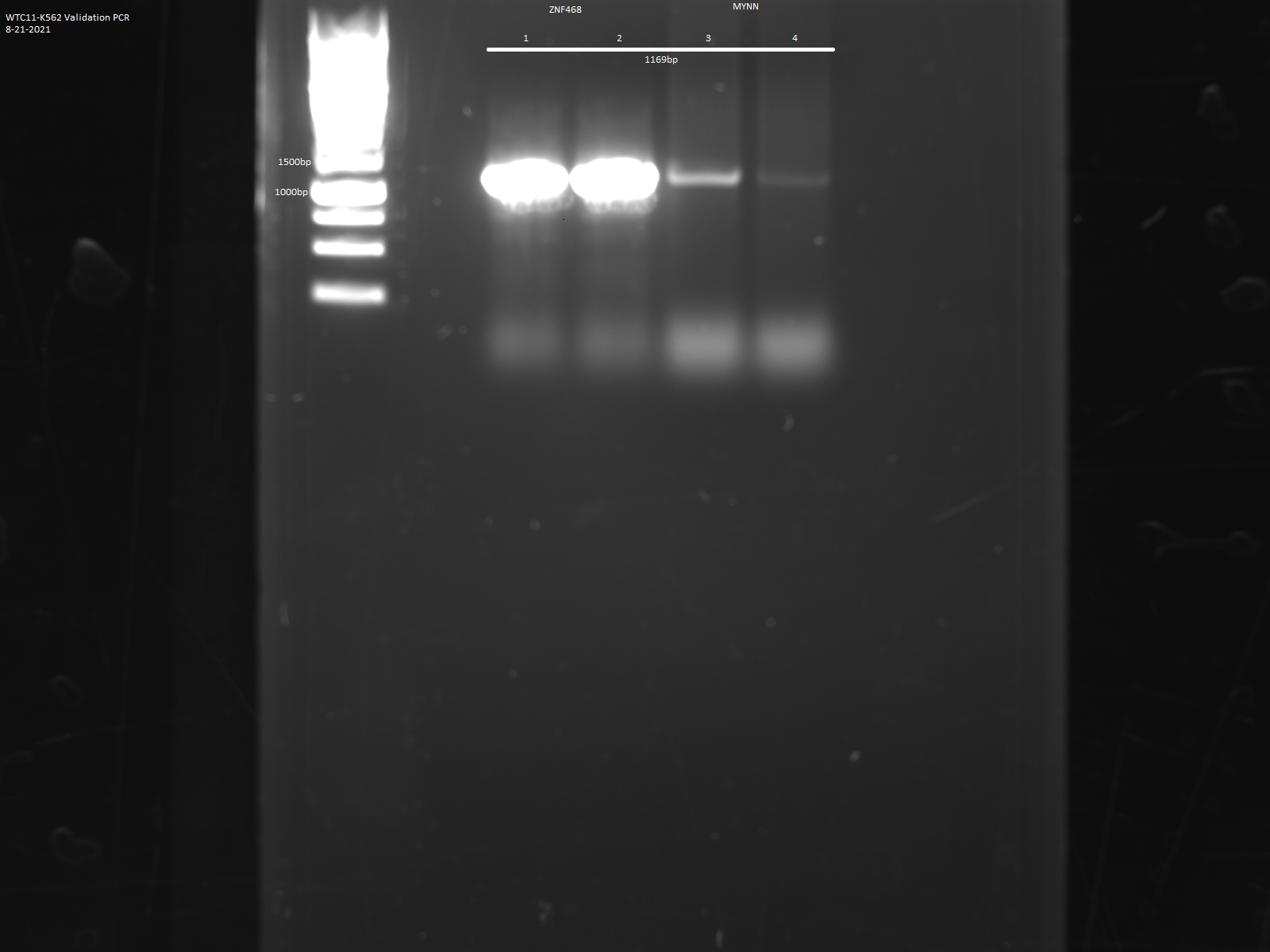
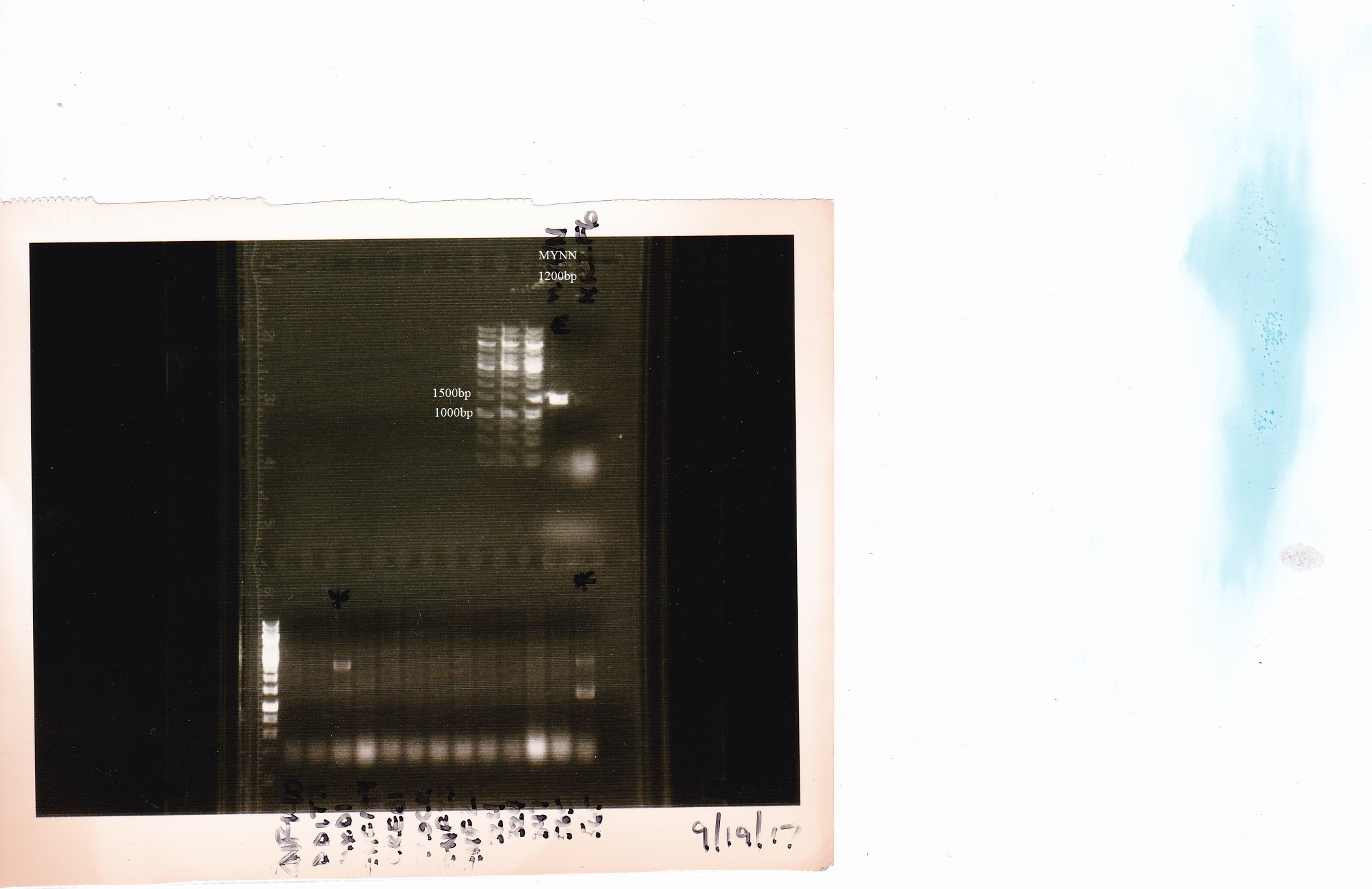
- CRISPR was used to insert a LAP tag downstream of the MYNN gene.
- Type
- insertion
- Tags
- eGFP — C-terminal
- Purpose
- tagging
Modification site
- Target
- MYNN-human
Modification method
- Method
- CRISPR
- Reagents
Documents
PCR analysis Characterization
- Lab
- Michael Snyder, Stanford
PCR analysis Characterization
- Lab
- Michael Snyder, Stanford